A new paper has been accepted at the 2024 IEEE International Conference on Bioinformatics and Biomedicine (BIBM)
Our recent study on “D-SarcNet: A Dual-stream Deep Learning Framework for Automatic Analysis of Sarcomere Structures in Fluorescently Labeled hiPSC-CMs” by Huyen Le et al. has been accepted for oral presentation at IEEE International Conference on Bioinformatics and Biomedicine 2024 (IEEE BIBM 2024). The IEEE International Conference on Bioinformatics and Biomedicine (BIBM) has established itself as the primer conference in bioinformatics and biomedicine. The IEEE BIBM 2024 provides a leading forum for disseminating the latest research in bioinformatics and health informatics. It brings together academic and industrial scientists from computer science, biology, chemistry, medicine, mathematics and statistics. This year, the IEE BIBM received more than 1660 research papers. In phase 2 paper submission, IEEE BIBM 2024 has received 965 paper submissions, and each paper was assigned to 5 Program Committee members for review. After the rigorous review process, 99% papers have received at least 3 reviews, and the conference has accepted 205 papers with an acceptance rate of 21%.

Figure 1. The 2024 Conference on Bioinformatics and Biomedicine will be held in Lisbon this December. The Politécnico de Portalegre, in collaboration with CERENA and IST is organizing this year’s conference, whose motto titles «Bridging The Research and Sectoral Shores To Tackle Digital Transition Opportunities». The event will bring together research communities on biomedical, health informatics, and life sciences.
In short, human-induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CMs) are a powerful tool in advancing cardiovascular research and clinical applications. The maturation of sarcomere organization in hiPSC-CMs is crucial, as it supports the contractile function and structural integrity of these cells. Traditional methods for assessing this maturation like manual annotation and feature extraction are labor-intensive, time-consuming, and unsuitable for high-throughput analysis. In this work, we propose D-SarcNet, a deep learning framework to automatically score sarcomere organization in fluorescently labeled hiPSC-CMs single-cell images, outperforming the current state-of-the-art without requiring the prior feature engineering process. We design a dual-stream framework architecture combining ConvNeXt and Swin Transformer for global and local feature extraction. Remarkably, we propose using FFT Power, deep learning-generated local patterns, and gradient magnitude as input for the second stream to provide the framework more information on the heterogeneity in sarcomeric organizational states. Extensive experiments and ablation studies demonstrate the advantages of the proposed framework over existing state-of-the-art methods and confirm the contribution of the proposed image-based representations in sarcomere analysis. The lack of progress in single-cell segmentation remains a limitation. In the AISC dataset, experts manually draw single-cell boundaries due to the lack of an accessible automated framework. This work should be deemed difficult because the single cells are overlap. Future work will focus on developing the hiPSC-CMs single-cell segmentation framework and exploring the potential of this framework in cardiac research pipeline.
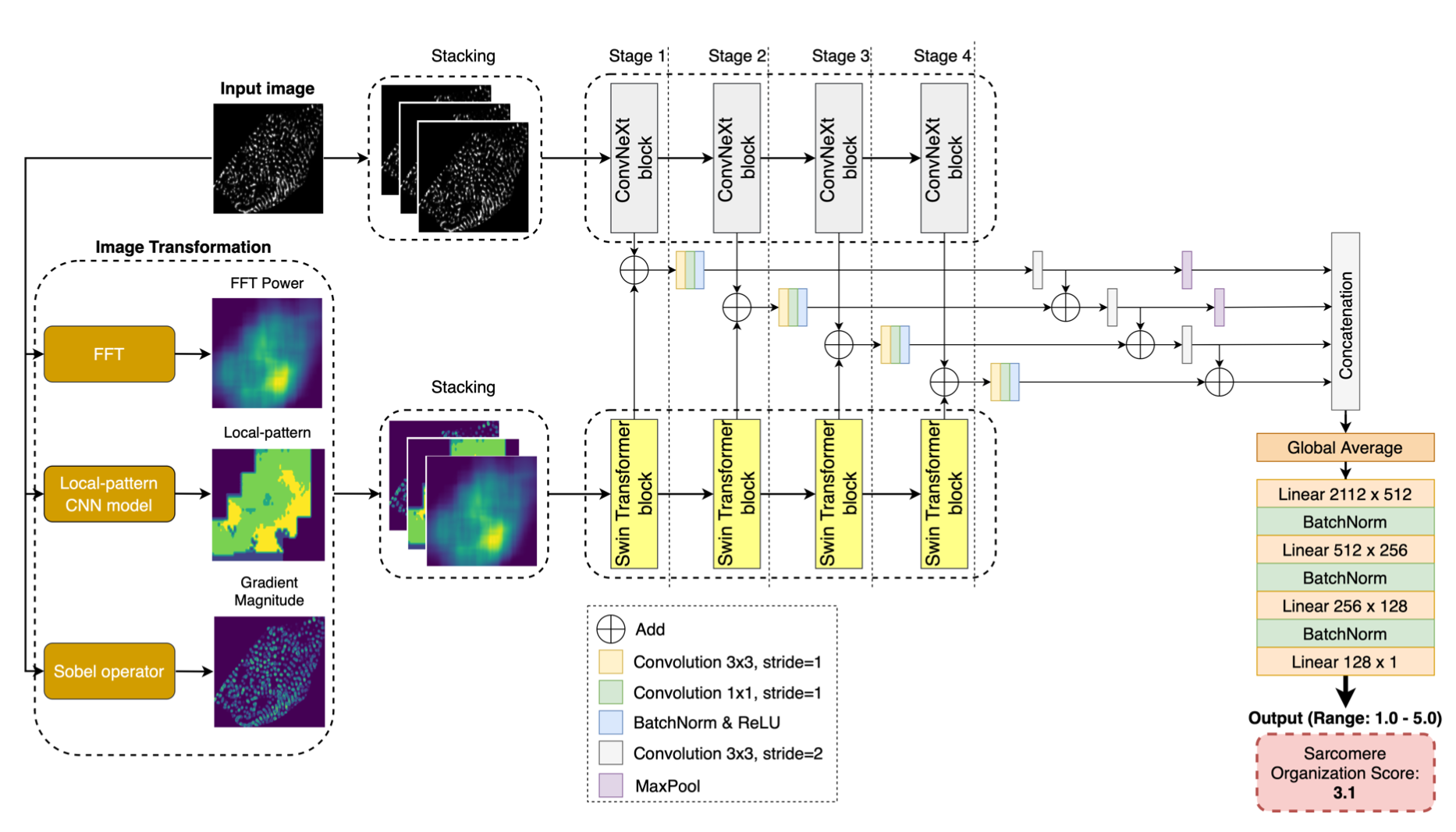
Figure 2. The D-SarcNet framework for scoring sarcomere structural organization in fluorescently labeled hiPSC-CM single-cell images consists of two primary streams. The first stream, ConvNeXt, processes raw images to extract global features directly from the original images. Simultaneously, the second stream, Swin Transformer, analyzes the corresponding three-channel image — created by stacking representations generated by FFT, the local pattern model, and the Sobel operator — to capture local features. A blocks-combined architecture then integrates feature maps from both streams across various scales to output a score of α-actinin-2 pattern structure on a scale from 1.0 to 5.0.
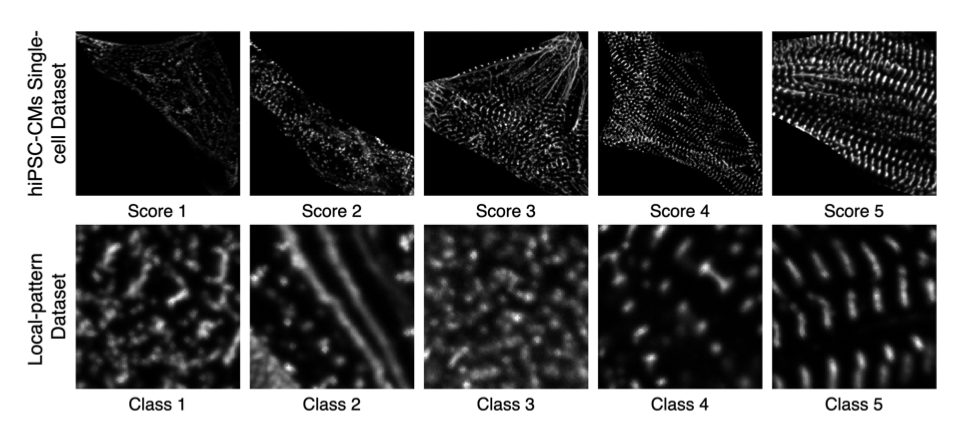

Figure 3. Representative examples of hiPSC-CMs single-cell images and local patterns for each category